Bandage is a program for visualising de novo assembly graphs. By displaying connections which are not present in the contigs file, Bandage opens up new possibilities for analysing de novo assemblies.
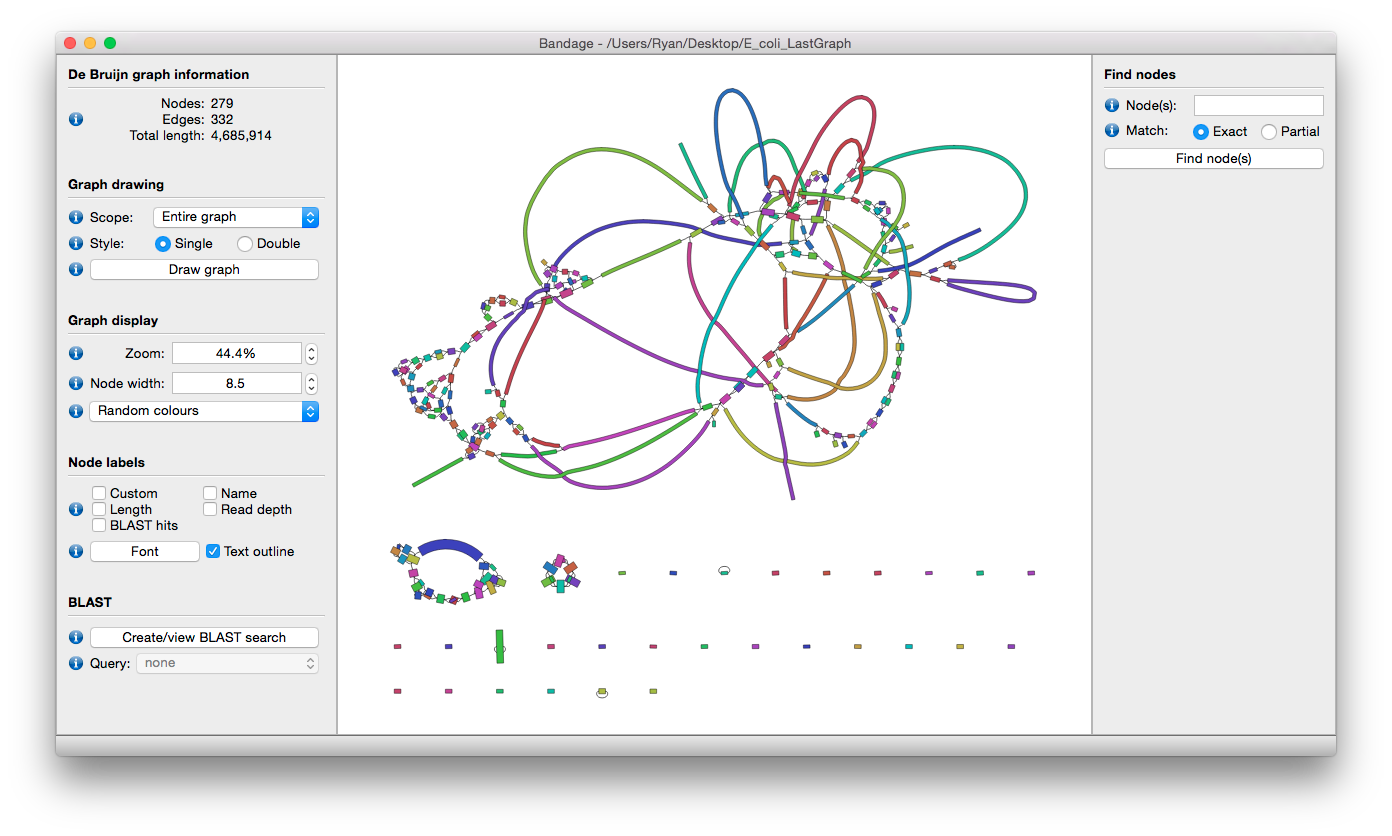
Features
- Load multiple assembly graph formats: LastGraph (Velvet), FASTG (SPAdes), Trinity.fasta, ASQG and GFA.
- Position nodes automatically with an efficient graph layout algorithm.
- Zoom, pan and rotate the view using either mouse or keyboard controls.
- Reposition and reshape nodes by clicking and dragging with the mouse.
- Configure graph scope: view the entire assembly graph or only a region of interest.
- Copy node sequences to the clipboard or save them to file.
- Colour nodes using built-in colour schemes or user-defined colours.
- Label nodes using node number, length, coverage or user-defined labels.
- Find nodes quickly in a large graph using node numbers.
- Specify the thickness of nodes and allow thickness to reflect the node's read depth.
- Define the relationship between the length of a node and the length of its sequence.
- Draw graph in single node style: each node and its reverse complements appear as a single object.
- Draw graph in double node style: nodes and their reverse complements appear as separate objects with arrow heads to indicate direction.
- Highlight and label specific sequences with integrated BLAST search.
- Automatically identify nodes contiguous with a node of interest.
- Call Bandage from the command line to specify settings, load graphs or generate images.
Uses
- Assess and compare the quality of assemblies quickly and visually.
- Identify problematic regions of an assembly.
- Resolve ambiguities in the graph to improve or complete de novo assemblies.
- Determine which other nodes have sequences that are contiguous with a node of interest.
- Annotate graph images to illustrate assembly features.
- Extract candidate sequences that extend through multiple nodes.
Installation
Bandage works equally well on OS X, Linux and Windows. Users are encouraged to download 64-bit binary executables using the links on the top-right of this page. No installation is necessary – just unzip and run. Windows and Mac binaries come packaged with all necessary libraries. The Linux binary has a couple of dependencies, specified in the zip file. More pre-built binaries are available on the GitHub 'Releases' page.
When first opening Bandage on a Mac, you may receive a warning stating that Bandage 'can't be opened because it is from an unidentified developer.' Right click on the file and select 'Open' to override this warning.
Alternatively, you can clone the source code from GitHub and build Bandage yourself. Instructions are on the GitHub page.
Sample data
Here are a couple of sample graph files you can download and view in Bandage:
- An E. coli assembly from Velvet. The reads and more information are available here.
- A small mouse transcriptome assembly from Trinity. This data comes with Trinity in the 'sample_data/test_Trinity_Assembly/' folder.
Documentation
Bandage's documentation is on the Bandage GitHub wiki.
Cite
If you use Bandage in your research, please cite the following publication:
